Lymphocyte Nuclear Biology
Our main goal is to unravel the nuclear events that drive peripheral activation of B lymphocytes. From genome-wide changes in epigenetics, chromatin architecture, and transcriptional activation, to recombination and hypermutation of antibody genes. Another major interest is to understand how deregulation of these mechanisms leads to B cell tumorigenesis. To achieve these goals, our team combines a wide range of cutting-edge technologies, including genome editing, nanoscopy, in situ Hi-C, cryo-EM, and bioinformatic tools.
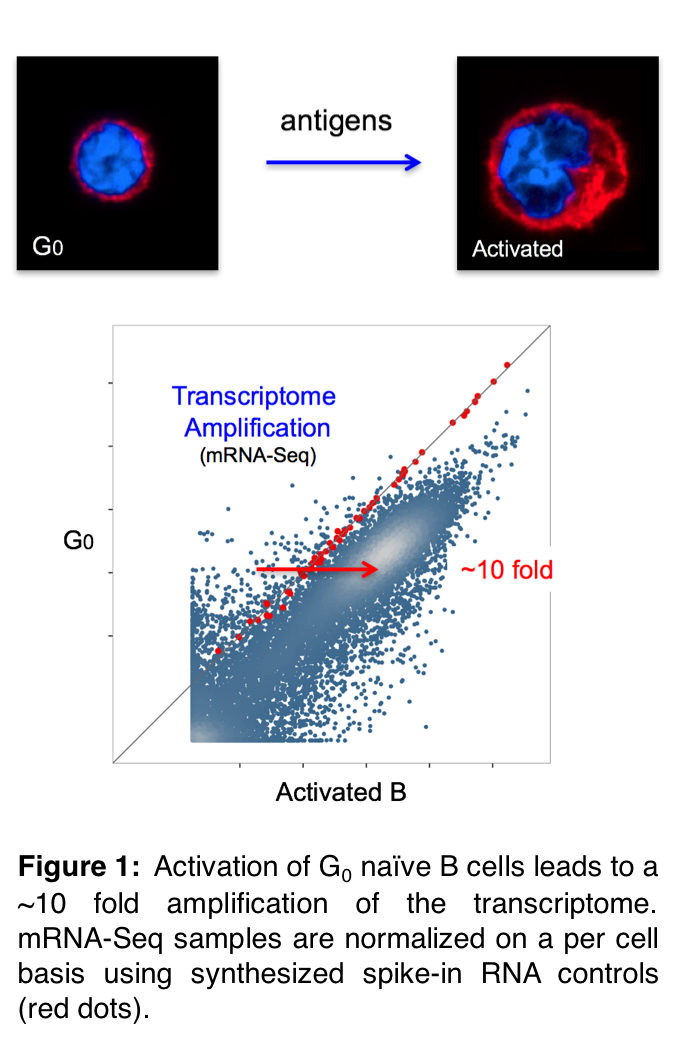 B lymphocytes in higher organisms are born in the bone marrow, where they expand and undergo V(D)J recombination to assemble their cell surface antibody receptors. The newly generated B cells then migrate to the periphery as quiescent G0 lymphocytes. In this metabolically inert state, B cells can live for up to 4 months. If during their lifespan they encounter foreign antigens, naïve B cells become activated and enter the cell cycle in less than 24h. How B lymphocytes are able to respond so rapidly to infection has been a mystery. In the past 5 years, we have resolved this puzzle by showing that the genome of naïve B cells is poised. RNA polymerases, transcription factors, chromatin remodelers, histone acetyl and methyltransferases, and nuclear architectural proteins are all recruited to G0 chromatin in a manner nearly identical to that seen in activated counterparts. In naïve cells, the preloaded enzymes only display basal activity until an optimal concentration of their catalytic substrates or cofactors is reached during activation. Taking transcription as an example, a comparative analysis between G0 and cycling cells shows that expression of all genes is proportionally amplified ~10 fold (Figure 1). We have dubbed this new biological phenomenon (so far unique to lymphocytes) transcriptome, epigenome, or architectural amplification.
B lymphocytes in higher organisms are born in the bone marrow, where they expand and undergo V(D)J recombination to assemble their cell surface antibody receptors. The newly generated B cells then migrate to the periphery as quiescent G0 lymphocytes. In this metabolically inert state, B cells can live for up to 4 months. If during their lifespan they encounter foreign antigens, naïve B cells become activated and enter the cell cycle in less than 24h. How B lymphocytes are able to respond so rapidly to infection has been a mystery. In the past 5 years, we have resolved this puzzle by showing that the genome of naïve B cells is poised. RNA polymerases, transcription factors, chromatin remodelers, histone acetyl and methyltransferases, and nuclear architectural proteins are all recruited to G0 chromatin in a manner nearly identical to that seen in activated counterparts. In naïve cells, the preloaded enzymes only display basal activity until an optimal concentration of their catalytic substrates or cofactors is reached during activation. Taking transcription as an example, a comparative analysis between G0 and cycling cells shows that expression of all genes is proportionally amplified ~10 fold (Figure 1). We have dubbed this new biological phenomenon (so far unique to lymphocytes) transcriptome, epigenome, or architectural amplification.
Activation of B cells also initiates a second round of antibody gene alterations. First, the Ch genes recombine to change the antibody isotype class, from IgM to IgG, IgE, or IgA. This process is known as class switch recombination and is mediated by the AID cytidine deaminase. In addition, the V domain of antibody genes is hypermutated, a process that coupled to selection enhances the affinity of antibodies for the immunogen. Like switch recombination, somatic hypermutation is driven by AID.
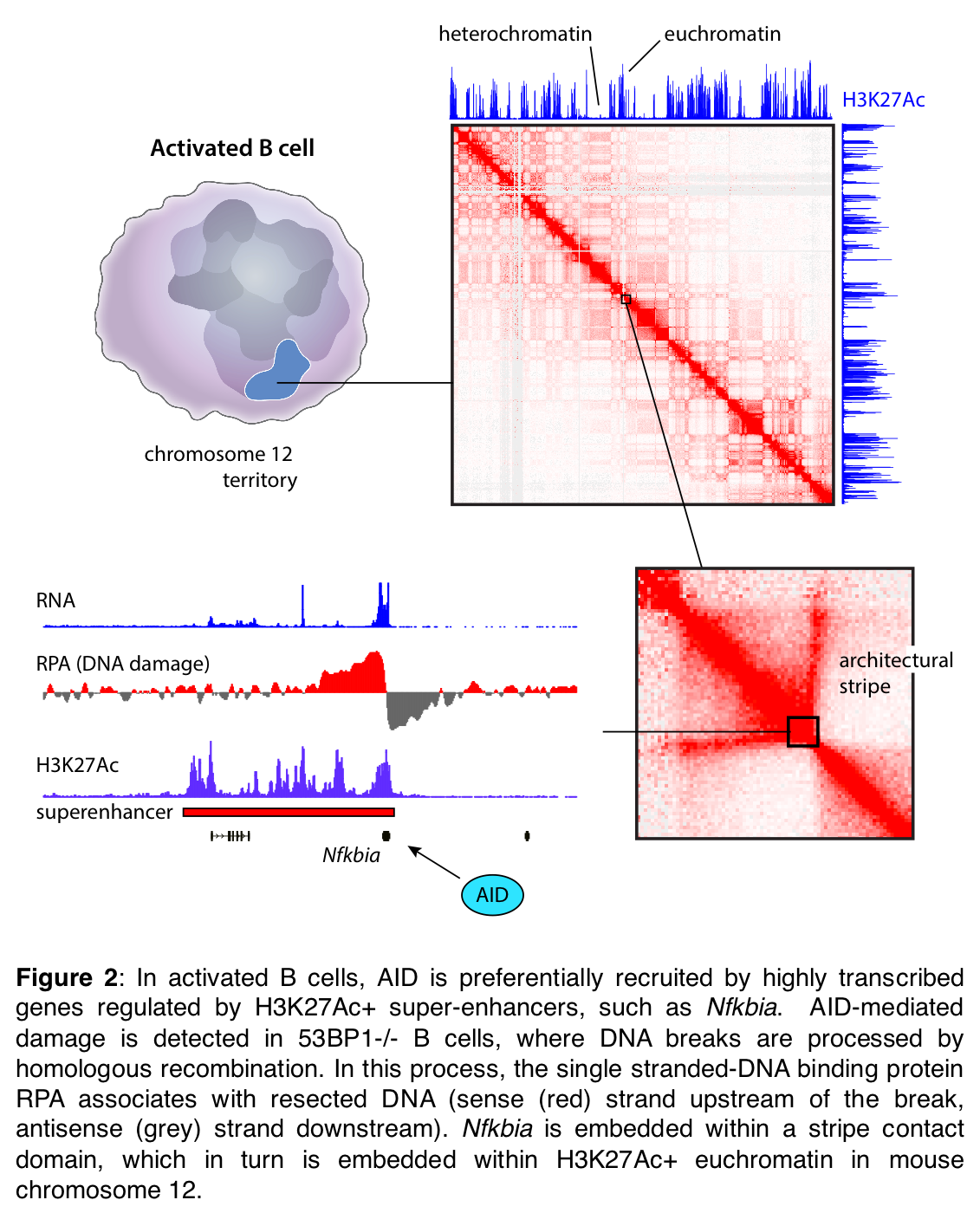
RAGs, and most particularly AID, can be promiscuous in nature, and the proclivity of B cells towards lymphomagenesis stems largely from AID-mediated lesions across the genome which can create chromosomal translocations between oncogenes (e.g. MYC, PIM1, MIR142) and antibody gene loci. In a series of key studies also in the past 5 years our laboratory has identified the molecular basis for AID promiscuity. We have found that AID is recruited by super-enhancers owing to their heightened accessibility, topological complexity, and superior transcriptional activity relative to other loci (Figure 2).
Current topics of interest in our laboratory include:
- The impact of nuclear architecture on gene expression.
- The role of the Mediator complex on B cell transcriptional regulation.
- The generation of new mouse models expressing antibodies of clinical interest